
M.D., Ph.D. Professor Naoki Watanabe
Cells change phenotypes such as migration, proliferation, differentiation and tissue formation sensing physical force from outer environment. We have proven that individual fluorescently-labeled proteins can be visualized in living cells. Using this technique, we elucidate dynamic interaction between cell signaling and cytoskeletal remodeling. For example, mDia1 and other formin homology proteins rapidly regenerate actin filaments in mechanically perturbed cells. At the cell leading edge, actin undergoes rapid polymerization and disassembly by a mechanism distinct from the widely-accepted ‘treadmilling’. We also apply our methods for elucidating the action of target-based drugs.
Research and Education
“Why not watching directly?” Cells dynamically change shape to undergo migration, proliferation, differentiation and tissue formation. The actin cytoskeleton is the major player in sensing and transducing mechanical signals as well as driving cell morphological changes, playing pivotal roles in development, tissue repair, immune cell migration, cancer cell invasion and so on. Because such dynamic processes operate in a complex manner even in a single cell, it has been difficult to precisely dissect individual molecular functions just by observing the phenotypes. Our invention of direct real-time viewing of individual molecules has brought a powerful tool to elucidate how complex biological systems operate in living cells. Using single-molecule imaging, we focus on;
- dynamic interaction between cell signaling and mechanotransduction.
- processive actin polymerization mechanisms of formin homology proteins.
- paradoxical signal activation by target-based cancerdrugs.
- Revolutionary multiplexed high-density labeling super-resolution microscopy IRIS.
Currently, our laboratory is most extensively working on our high-density labeling multi-target super-resolution microscopy IRIS to extend its applications to animal disease models.
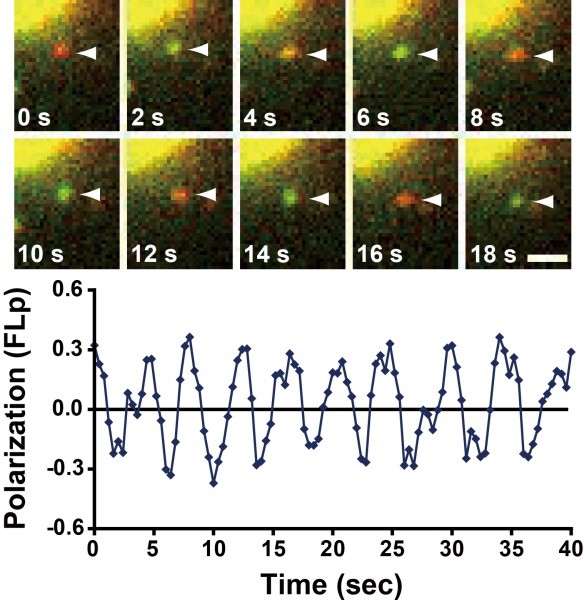 Figure 1. Helical actin polymerization by mDia1 visualized by single-molecule fluorescence polarization(Science 2011)
Figure 1. Helical actin polymerization by mDia1 visualized by single-molecule fluorescence polarization(Science 2011)
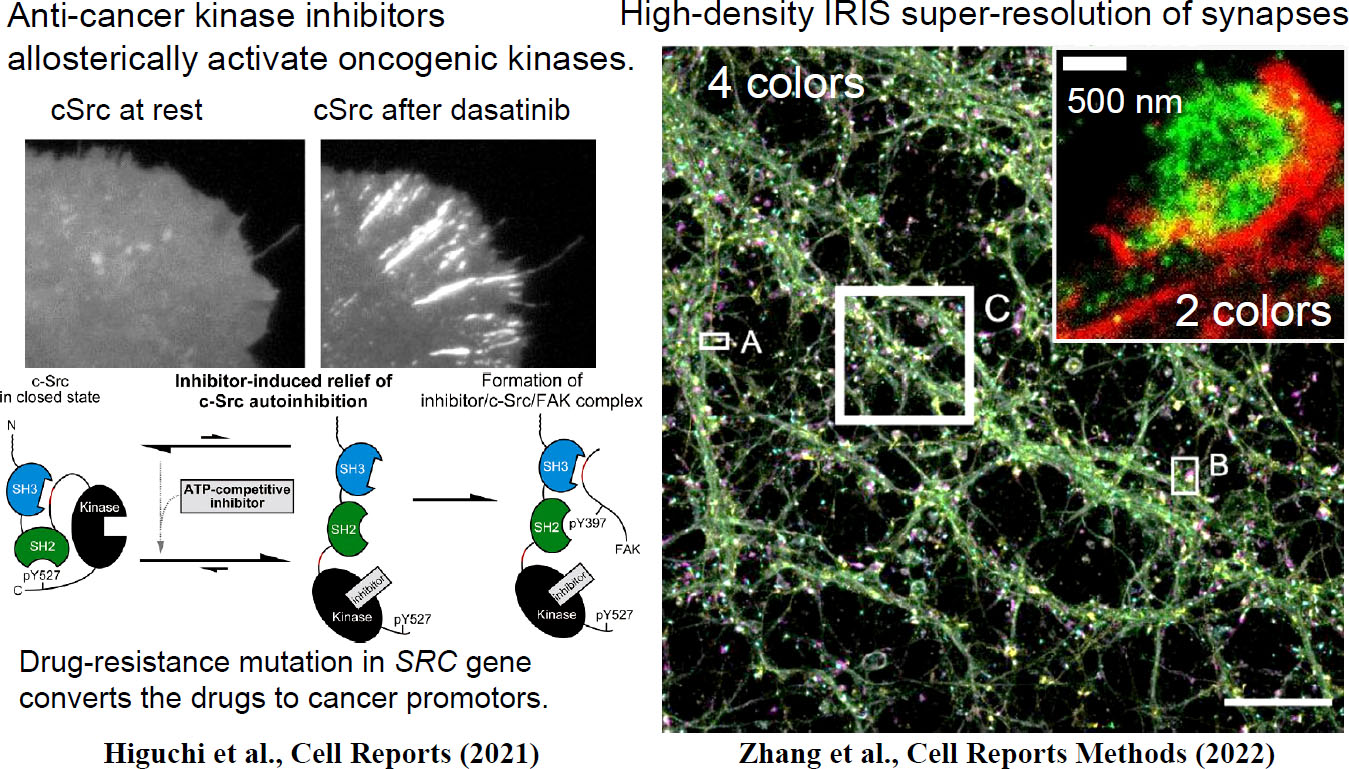 Figure 2
Figure 2
Recent Publications
- Mizuno, H., Tanaka, K., Yamashiro, S., Narita, A. and Watanabe, N. (2018) Helical rotation of the diaphanous-related formin mDia1 generates actin filaments resistant to cofilin. Proc. Natl. Acad. Sci. USA 115, E5000-E5007.
- Kiuchi, T., Higuchi, M., Takamura, A., Maruoka, M. and Watanabe, N. (2015) Multitarget super-resolution microscopy with high-density labeling by exchangeable probes. Nat. Methods 12, 743-746.
- Higashida, C., Kiuchi, T., Akiba, Y., Mizuno, H., Maruoka, M., Narumiya, S., Mizuno, K. and Watanabe, N. (2013) F- and G-actin homeostasis regulates mechanosensitive actin nucleation by formins. Nat. Cell Biol. 15, 395-405.
- Mizuno, H., Higashida, C., Yuan, Y., Ishizaki, T., Narumiya, S. and Watanabe, N. (2011) Rotational movement of the formin mDia1 along the double helical strand of an actin filament. Science 331, 80-83.
- Higashida C. Miyoshi T., Fujita A., Oceguera-Yanez F., Monypenny J., Andou Y., Narumiya S. and Watanabe N. (2004) Actin polymerization-driven molecular movement of mDia1 in living cells. Science 303, 2007-2010.
Laboratory
Professor: Naoki Watanabe
Associate Professor: Tai Kiuchi
Lecturer: Sawako Yamashiro
Assistant Professor: Akitoshi Miyamoto
TEL: +81-75-753-4396
FAX: +81-75-753-4394
e-mail: watanabe.naoki.4v <at> kyoto-u.ac.jp
URL: http://www.pharm2.med.kyoto-u.ac.jp/2_index.html
