
Ph.D. Professor Yasushi Okuno
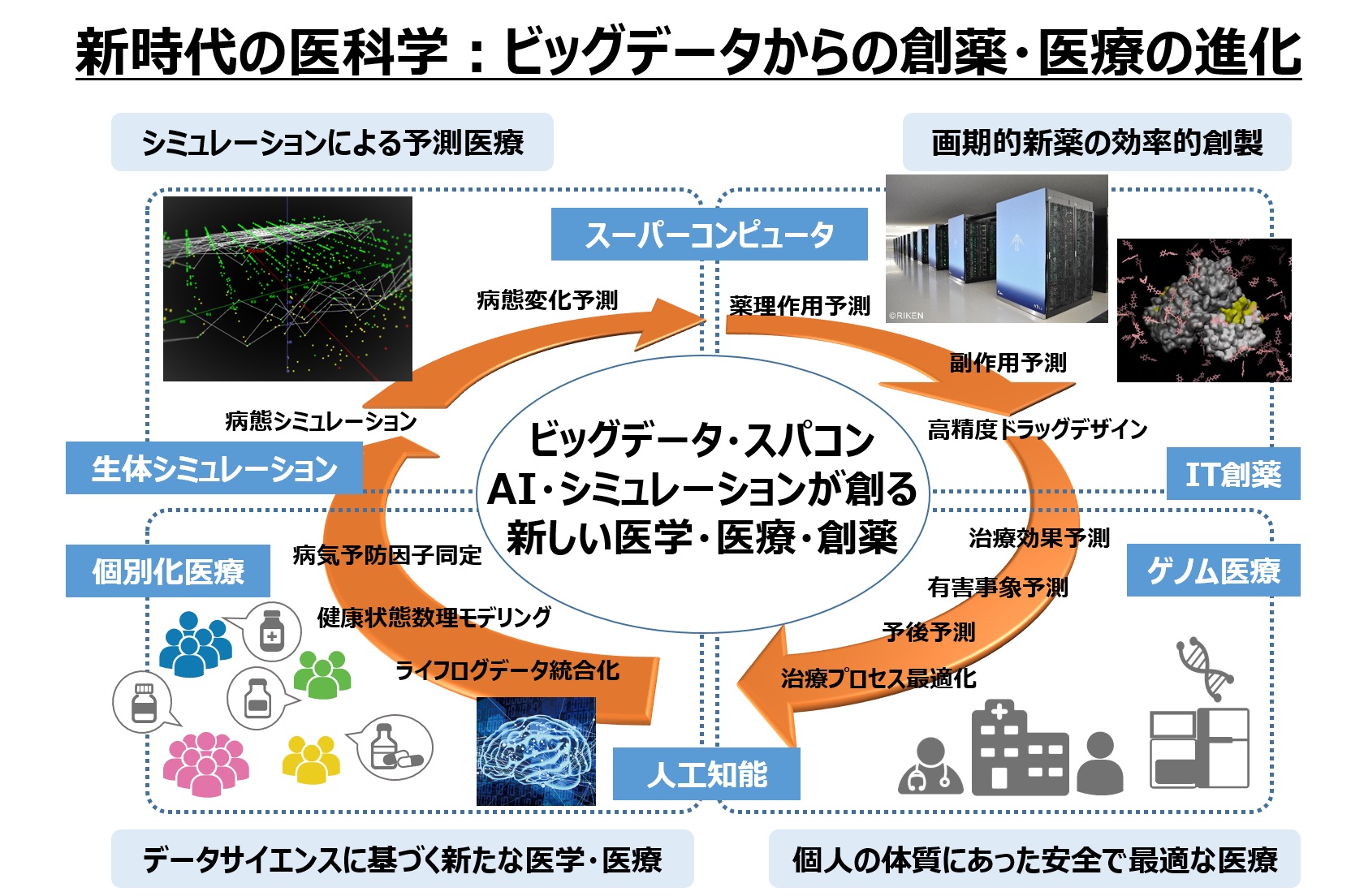
In addition to experimentally-driven and theoretically-driven science, two new waves of science are globally emerging that focus on simulation science and data-centric science. Our laboratory is developing new methods for medical big data analysis and medical AI using actual clinical data from the Kyoto University Hospital, as well as new methods for simulation drug discovery and AI drug discovery by using the supercomputer ”Fugaku”. Through these developments we aim to open the way to simulation science and data science for practical medical care and applied drug discovery.
Research and Education
Medical big data analysis and medical AI:
In collaboration with the Kyoto University Hospital Cancer Center Biobank & Informatics for Cancer Project, which has established a protocol for longitudinal recording of clinical data as well as collection and storage of blood samples, the laboratory aims to develop new methods for integration and analysis of various types of biological data, and in doing such, provide frameworks that enable clinical data to spur the creation of translational clinical informatics, data-driven personalized medicine, and development of healthcare. More specifically, we are using AI and simulation techniques to create new methods for systematic, rational inference about each individual cancer patient’s physical state, efficacy of treatment, and prediction of drug adverse events. In relation to such data, we are additionally developing methods for novel biomarker identification and drug target discovery.
Simulation and AI drug discovery:
In recent years, the pharmaceutical industry has been faced with a crucial problem of how to efficiently develop new drugs while reducing development costs, and there is high expectation for the use of computers to perform “in-silico drug discovery”. Notably, with the new availability of the supercomputer ”Fugaku” has come the blossoming of computational science and informatics for drug discovery. Our laboratory has established a consortium of pharmaceutical companies, IT companies, and academic labs that is developing drug discovery computational technologies using AI and the supercomputer ”Fugaku”.
Recent Publications
- Nakamura K, Kojima R, Uchino E, Ono K, Yanagita M, Murashita K, Itoh K, Nakaji S, Okuno Y. Health improvement framework for actionable treatment planning using a surrogate Bayesian model. Nat Commun. 2021;12(1):3088. doi:10.1038/s41467-021-23319-1
- Araki M, Matsumoto S, Bekker GJ, Isaka Y, Sagae Y, Kamiya N, Okuno Y. Exploring ligand binding pathways on proteins using hypersound-accelerated molecular dynamics. Nat Commun. 2021;12(1):2793. doi:10.1038/s41467-021-23157-1
- Matsumoto S, Ishida S, Araki M, Kato T, Terayama K, Okuno Y. Extraction of protein dynamics information from cryo-EM maps using deep learning. Nat Mach Intell. 2021;3:153–160. doi:10.1038/s42256-020-00290-y
- Mizuta H, Okada K, Araki M, Adachi J, Takemoto A, Kutkowska J, Maruyama K, Yanagitani N, Oh-Hara T, Watanabe K, Tamai K, Friboulet L, Katayama K, Ma B, Sasakura Y, Sagae Y, Kukimoto-Niino M, Shirouzu M, Takagi S, Simizu S, Nishio M, Okuno Y, Fujita N, Katayama R. Gilteritinib overcomes lorlatinib resistance in ALK-rearranged cancer. Nat Commun. 2021;12(1):1261. doi:10.1038/s41467-021-21396-w
- Saito Y, Koya J, Araki M, Kogure Y, Shingaki S, Tabata M, McClure MB, Yoshifuji K, Matsumoto S, Isaka Y, Tanaka H, Kanai T, Miyano S, Shiraishi Y, Okuno Y, Kataoka K. Landscape and function of multiple mutations within individual oncogenes. Nature. 2020;582(7810):95–99. doi:10.1038/s41586-020-2175-2
Laboratory
Professor:
Yasushi Okuno
Associate Professor:
Ryosuke Kojima
Associate Professor:
Mitsugu Araki, Shigeyuki Matsumoto, Yohei Mineharu, Takao Otsuka, Eiichiro Uchino
Senior Lecturer:
Hirohiko Kohjitani
Assistant Professor:
Minoru Sakuragi
Assistant Professor:
Yuji Okamoto, Mai Nakazawa, Itsuki Maeda
TEL/FAX: 075-751-3920
URL: http://clinfo.med.kyoto-u.ac.jp
